Python Starter Kit
Overview
This module is meant to be a starter kit for learning the basics of python. The goal is to give the reader the basics such that they can write basic scripts, and understand many major concepts. It is not comprehensive, and more thorough learning experience would not consist of a single module – but rather many modules. There is some prior content in other modules that may be helpful for the user. After the primary material is given, there are some examples for the user to test.
This module should not be your primary resource. One big piece of advice that is always there – one learns when they hit an error. A great way to solve an error is to just Google the error.
Finally, these modules are in development. If you see an error, please feel free to email davidwcr@usc.edu or leave a comment.
Learning environment
Like R, and most other languages, Python is developed and implemented in a variety of tools. One common way is to simply create scripts within the command-line. This would be done by opening up the command-line interface (typically called “Terminal” in a MacOS environment), and creating a file with your favorite text editor. Ideally, that is vim or emacs and not nano. There are other ways using graphical tools, such as Sublime3, and text-editors, then simply save the file. Most linux/unix based platforms include a version of python, but in some cases, they must be installed. On a Mac, these are installed by going to the App Store and installing XCode or using Brew. We cover this in a different module.
Overall, this module was developed originally for the command-line, and is being ported over to Jupiter. We do still have command-line instructions, and at some level, its good to see those since so many recipes presume the command-line. That said, it should always be straight forward to adapt these to Jupytor.
You are welcome to install Jupyter on your computer, but we will generally just presume the Browser based version, which is here.
High-level: What is Python?
Python is an interpreted, general-purpose programming language that generally is considered easy to read. Interpreted means that one does not need to compile the code to machine language separately, as is the case with C or Fortran. Python has an extensive set of libraries that can be added, given it considerable utility. You may hear of Python 2 and Python 3. Well as of January 1st, 2020, Python 2 is no longer supported. There are certain parts of the syntax of Python 2 that do not work in Python 3, and thus it is not considered backward compatible. Some important characteristics of Python are that its object-oriented. This is a programming paradigm based on the concept of objects containing data (like a box), and some methods you can do to the box. We might of person as an object, with person.name as where their name is stored. The person object comes with some functions, and one can do. For example, person.name is a str type, and thus every function associated with str becomes inherited. Thus, as we will see later, we have built-in ways to do things like make uppercase person.name.upper()where upper() is the way we make upper case.
Getting Used to Our Development Environment
JupyterLab and Notebooks
JupyterLab and Jupyter Notebooks are a graphic interface for developing and writing Python code – and can be used for many other languages such as R, scala etc. A notebook usually means a single analysis or collection of scripts/code. The JupyterLab is a place for storing and managing many Notebooks.
Using Browser-based Jupyter Notebook
You are welcome to install Jupyter on your computer, but we will generally just presume the Browser based version, which is here.

More Advanced Jupyter Lab!
Jupyter Lab provides more functionality, and eventually is where you want to go. In many parts of the tutorial, we just use the notebook. However, if you want full functionality and saving, you do want to move to installing locally Jupyter Lab. There is a version that can be run within your browser.
A snapshot is shown below. You can see in this view, you can see all files, you can edit and create what are called markdowns in the middle frame, and you have rendered markdowns in the right frame. We have discussed markdowns before, but basically they are a way of putting code and text together in a way that looks like a web page.

The code is run in cells by pressing the forward arrow after clicking on the cell.
You can see there are a lot of different types of files that can be created, and we discuss these in other modules. For this exercise, we largely just focus on the notebook

A few quick notes on the learning environment.
The default setup comes with some basic instruction of the notebook and Python. I recommend clicking on “Python” – beyond plain python.
This opens up a Markdown file. We discussed Markdowns previously, and the concept is fairly similar. There are areas where you put text and commentary, there are areas where code resides, and there are places where the output appears.


Running Python code
Above runs the code for print(“Hi”). Now if we were running in the command-line, we would put that within a file – ideally ending in py, like hello.py, and add a python call to the top. The call or shebang would be typically!#/usr/bin/python where that points to the full path of python in your computer. One would need to change the permission typically to be executable, such as chmod 755 hello.py. We cover these command-line concepts in an earlier module, and if you have questions, its best to go to that section.
In Juypiter, its a bit simpler. To run a line of code, simply click first on the code block you wish to run, and then click “Run” at the top.

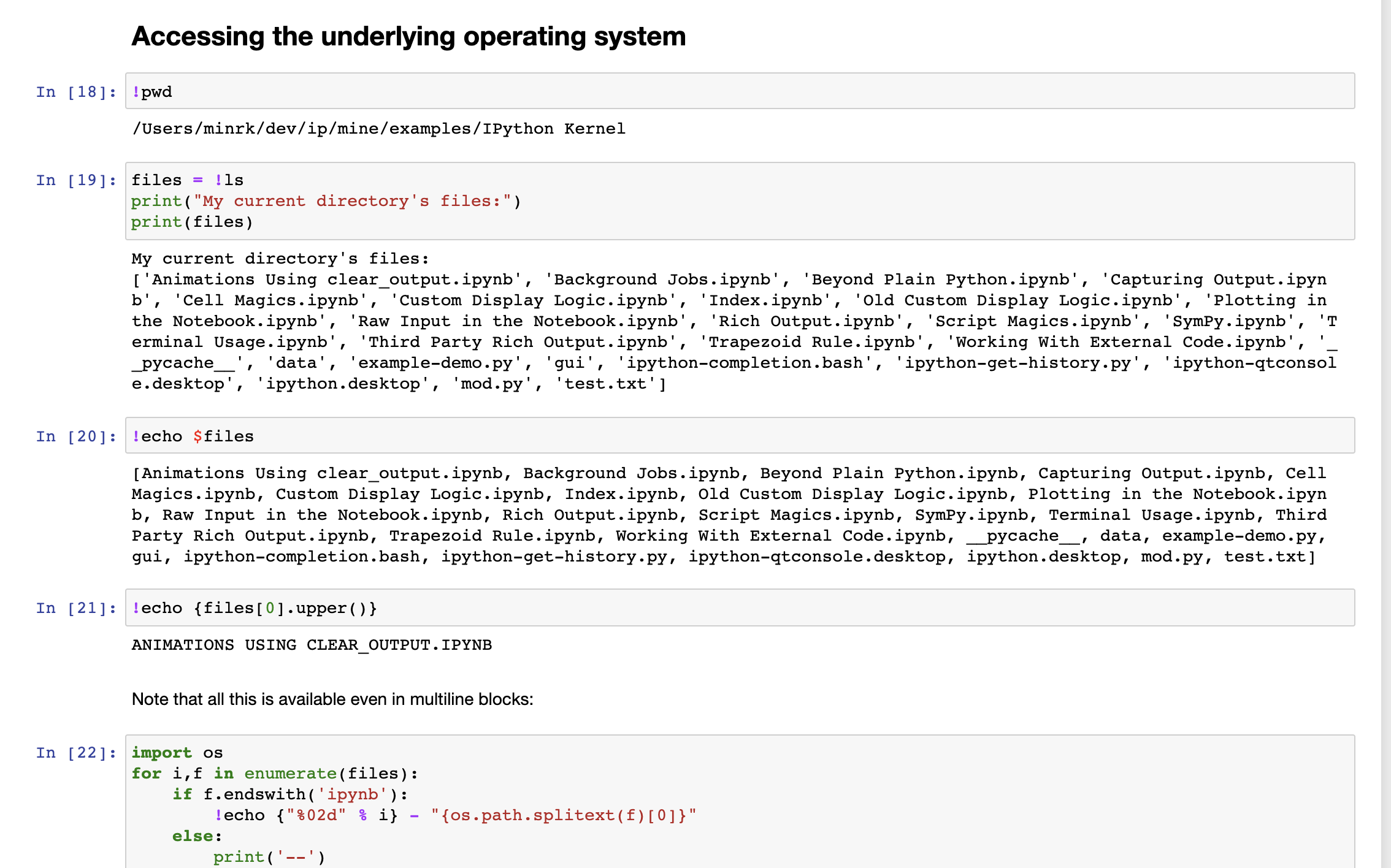
The nice thing about Jupyter is that we do have access to an underlying command-line and filesystem. These are described in the IJupyter.
Accessing the underlying operating system

Building a new notebook
Once we are comfortable, lets go ahead and create a new notebook that we will use for these exercises.

This starts a binder where we can begin editing and creating code. To create commentary or notes, you’ll need to switch from code to header.
![]()
![]()
We then create a header using standard Markdown nomenclature.

To see it in a more rendered view, click Run.
A few more magic things in Jupyter
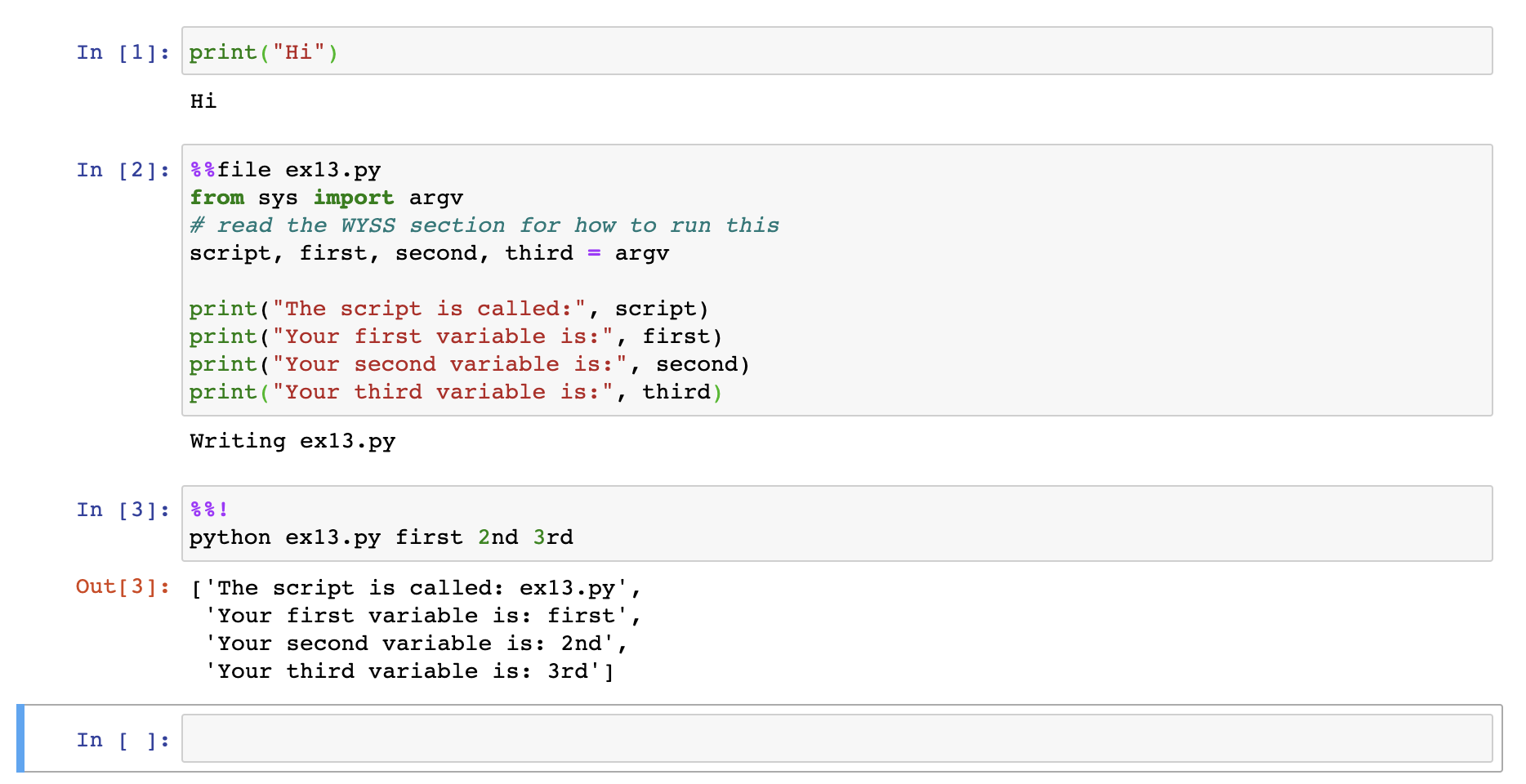
Often we do see programs that are meant to be at the command line, and they take command-line arguments. For these, Jupyter offers magic variables that provide a way to mimic these. Below we see an example creating a program ex13.py and then running in a later part of our notebook.

Some resources for learning a few more details
Interactive Learning
Please start an interactive environment.
We are going to have you create a python script in your browser, and then run it. It will print “hello” and tell you the name of your script.
Basics Hello, World
There are times when one wishes to edit or make more robust scripts that are simply beyond the capabilities of bash. Python is a highly useful language and not the focus of this course. We do take one module to teach some very basics. There are excellent courses on python and these are encouraged recognizing that not all bioinformaticians use and know python. Python is a dynamic, interpreted (bytecode-compiled) language. There are no type declarations of variables, parameters, functions, or methods in source code.
Python source files use the “.py” extension and are called modules or scripts. We will create a script called hello.py. The first line in our script `%file hello.py` is specific to Jupyter. It basically stores the text in subsequent lines in a new file called hello.py. When run it, the contents after this line are in the file hello.py
The second line imports libraries. There is a library called argv that allows us to access information from when the program was run. Sometimes we want options with programs. An example of this would be the program ls which normally simply lists the files in the directory. Sometimes, we want more information and we type ls -l . In this example, the -l is an argument passed into the program ls. The argv is the library you want if you are going to access these.
With a Python script hello.py, the easiest way to run it is with the shell command `python hello.py Alice” which calls the Python interpreter to execute the code in hello.py, passing it the command line argument “Alice”.
%%file hello.py
from sys import argv
script, name = argv
print ('Hello there', name,'. You ran: ',script)
We run this in a new cell
%%!python hello.py david
['Hello there david . You ran: hello.py']
The way this works is that you want to paste or type in the code within the cell from the notebook, then press Run. See this picture for an example:
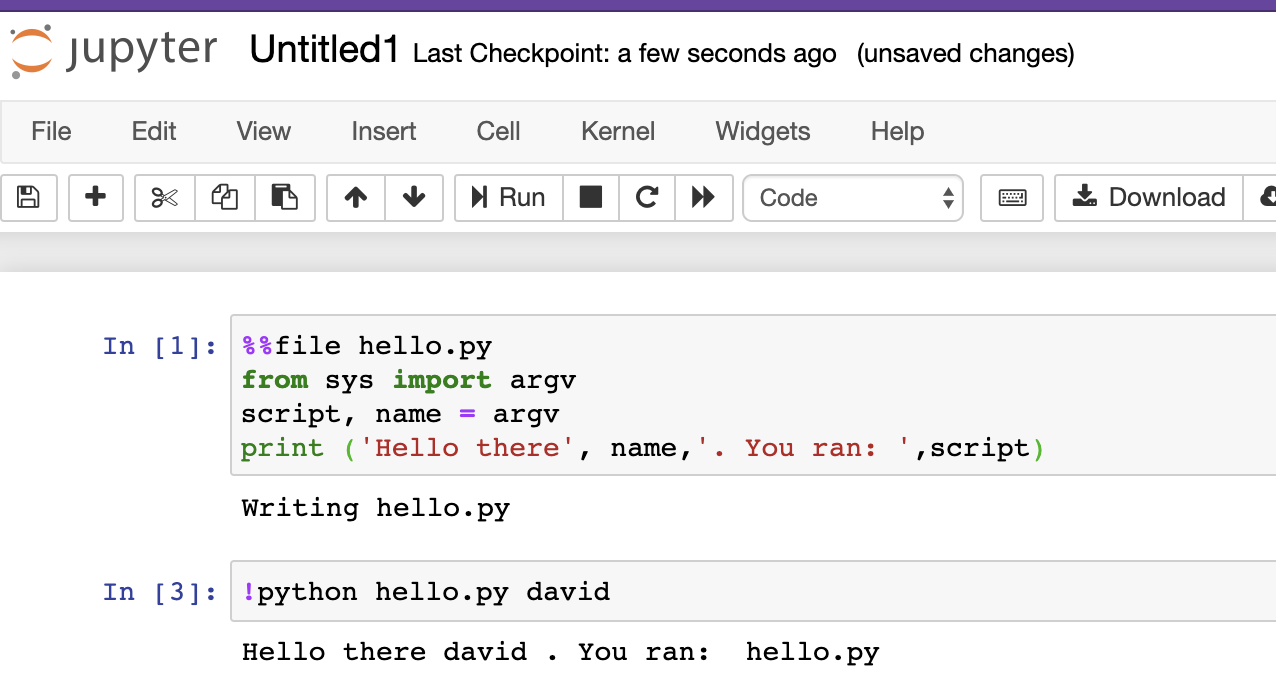

You can see the first cell created the file hello.py, and then second line ran python, feeding it the script hello.py. In this example, we also gave an additional command-line argument david, which was used by the script. In the script, name received the text “david”.
Now, if we didn’t put a name in, we’d see an error:
['Traceback (most recent call last):', ' File "hello.py", line 2, in <module>', ' script, name = argv', 'ValueError: not enough values to unpack (expected 2, got 1)']
Later we will talk about how to check if the user put in a name, rather than crashing
Python Syntax
The first thing to learn with a new language is a little about the syntax. Those are the rules of the language.
Python should have extension py when in unix. Python does require a first or last line, but typically begins with #!/usr/bin/python unless you wish to run it as python file.py. If this line isn’t there, you need to run it by typing python myscript.py.
Unlike many languages, there are no braces or semicolons (to end statements). Blocks are identified by having the same indention – indention matters.
%%file example1.py
#!/usr/bin/python
bool = True
name = "Craig"
age = 26
pi = 3.14159
print(name + ' is ' + str(age) + ' years old.')
print('Print requires parenthesis')
print("and single or double quotes")
print("Newlines can be escaped like\nthis.")
print("This text will be printed"),
print("on one line becaue of the comma.")
print ("Did you know: \n pi is ",pi,"?")
name = input("Enter your name: ")
a = int(input("Enter a number: "))
print(name + "'s number is " + str(a))
print("Lets make a and b 5, and then add 4.")
a = b = 5
a = a + 4
print (' a is now ', a, ', and b is ',b)
To run this example, we would need to make the permission executable and thus:
!python example1.py
The output is:
Craig is 26 years old.
Print requires parenthesis
and single or double quotes
Newlines can be escaped like
this.
This text will be printed
on one line becaue of the comma.
Did you know:
pi is 3.14159 ?
Enter your name: davd
Enter a number: 54
davd's number is 54
Lets make a and b the same, and then add.
a is now 9 , and b is 5
Indentation
One unusual Python feature is that the whitespace indentation of a piece of code affects its meaning. A logical block of statements such as the ones that make up a function should all have the same indentation, set in from the indentation of their parent function or “if” or whatever. If one of the lines in a group has a different indentation, it is flagged as a syntax error.
Python’s use of whitespace feels a little strange at first, but it’s logical and I found I got used to it very quickly. Avoid using TABs as they greatly complicate the indentation scheme (not to mention TABs may mean different things on different platforms). Set your editor to insert spaces instead of TABs for Python code.
A common question beginners ask is, “How many spaces should I indent?” According to the official Python style guide (PEP 8), you should indent with 4 spaces.
Comments
Putting notes to yourself and others, is well an essential part of programming. In Python, this is done with the hash # symbol, where everything that follows that line is ignored.
Variables and datatypes
Variables can be boolean, integer, long, float,string, list or an object. Variables are local – within their own function or class unless otherwise defined as “global”.
Since Python variables don’t have any type spelled out in the source code, it’s extra helpful to give meaningful names to your variables to remind yourself of what’s going on. So use name if it’s a single name, and names if it’s a list of names, and tuples if it’s a list of tuples. Many basic Python errors result from forgetting what type of value is in each variable, so use your variable names (all you have really) to help keep things straight.
As you can guess, keywords like print and while cannot be used as variable names — you’ll get a syntax error if you do. However, be careful not to use built-ins as variable names. For example, while ‘str’ and ‘list’ may seem like good names, you’d be overriding those system variables. Built-ins are not keywords and thus, are susceptible to inadvertent use by new Python developers.
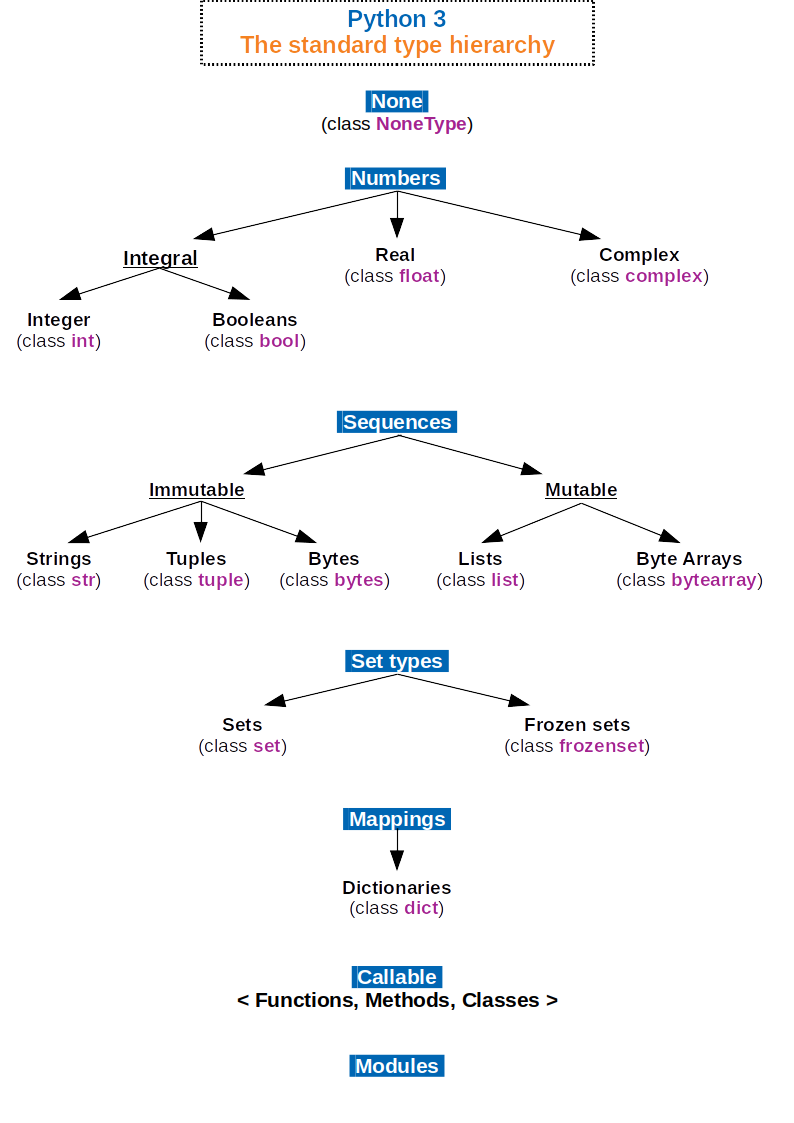

Datatypes: Strings
Python has a built-in string class named str with many handy features. String types can be enclosed by either double or single quotes, and single quotes are more often used. Backslash escapes work the usual way within both single and double quoted literals — e.g. n "hello". A double quoted string literal can contain single quotes without any issue (e.g. "hello") and likewise single quoted string can contain double quotes"hello, 'he said'". A string literal can span multiple lines, but there must be a backslash at the end of each line to escape the newline. String literals inside triple quotes, """ or "", can span multiple lines of text.
Python strings are “immutable” which means they cannot be changed after they are created. Since strings can’t be changed, we construct new strings as we go to represent computed values. For example, the expression ("hello" + "there") takes in the 2 strings hello and there and builds a new string hellothere.
Characters in a string can be accessed using the standard [ ] syntax, and like Java and C++, Python uses zero-based indexing, so if str is hello str[1] is e. If the index is out of bounds for the string, Python raises an error. The Python style (unlike Perl) is to halt if it can’t tell what to do, rather than just make up a default value. The handy “slice” syntax (below) also works to extract any substring from a string. The len(string) function returns the length of a string. The [ ] syntax and the len() function actually work on any sequence type — strings, lists, etc.. Python tries to make its operations work consistently across different types. Python newbie gotcha: don’t use len as a variable name to avoid blocking out the len() function. The + operator can concatenate two strings. Notice in the code below that variables are not pre-declared — just assign to them and go.
Strings can be specified using quotes or double quotes:
#!/usr/bin/python
a = "Bioinformatics"
print ("The value of a is \t",a) # I snuck in a tab using \t
s="Bioinformatics is not too difficult to learn"
# String functions as char arrays
x = s[0:5]
print ("The value of x is ", x)
x = s[:8]
print ("The value of x is now ",x)
x = s[11:]+", unless I make errors"
print ("The value of x is longer with ", x)
x = len(s)
#print "Try: The value of x is " + x
print ("Try: The length of s (",s,") is ",x)
pi = 3.14 ##text = 'The value of pi is ' + pi ## NO, does not work text = 'The value of pi is ' + str(pi) ## yes print(text)
For numbers, the standard operators, +, /, * work in the usual way. There is no ++ operator, but +=, -=, etc. work. If you want integer division, it is most correct to use 2 slashes — e.g. 6 // 5 is 1 (previous to python 3000, a single / does int division with int anyway, but moving forward // is the preferred way to indicate that you want int division.)
The “print” operator prints out one or more python items followed by a newline (leave a trailing comma at the end of the items to inhibit the newline). A “raw” string literal is prefixed by an r and passes all the chars through without special treatment of backslashes, so r'xnx' evaluates to the length-4 string xnx. A u prefix allows you to write a Unicode string literal (Python has lots of other Unicode support features — see the docs below).
#!/usr/bin/python raw = r'thist and that' print (raw) ## thistn and that multi = """It was the best of times. It was the worst of times.""" print (multi)
thistn and that It was the best of times. It was the worst of times.
String Methods
Here are some of the most common string methods. A method is like a function, but it runs “on” an object. If the variable s is a string, then the code s.lower() runs the lower() method on that string object and returns the result (this idea of a method running on an object is one of the basic ideas that make up Object Oriented Programming, OOP). Here are some of the most common string methods:
s.lower(),s.upper()— returns the lowercase or uppercase version of the strings.strip()— returns a string with whitespace removed from the start and ends.isalpha()/s.isdigit()/s.isspace()… — tests if all the string chars are in the various character classess.startswith('other'),s.endswith('other')— tests if the string starts or ends with the given other strings.find('other')— searches for the given other string (not a regular expression) within s, and returns the first index where it begins or -1 if not founds.replace(‘old’, ‘new')— returns a string where all occurrences of ‘old’ have been replaced by ‘new’s.split(‘delim’)— returns a list of substrings separated by the given delimiter. The delimiter is not a regular expression, it’s just text.aaa,bbb,ccc.split(',')-> [‘aaa’, ‘bbb’, ‘ccc’]. As a convenient special case s.split() (with no arguments) splits on all whitespace chars.s.join(list)— opposite of split(), joins the elements in the given list together using the string as the delimiter. e.g. ‘—‘.join([‘aaa’, ‘bbb’, ‘ccc’]) -> aaa—bbb—ccc
A google search for “python str” should lead you to the official python.org string methods which lists all the str methods.
Python does not have a separate character type. Instead, an expression like s[8] returns a string-length-1 containing the character. With that string-length-1, the operators ==, <=, … all work as you would expect, so mostly you don’t need to know that Python does not have a separate scalar “char” type.
String Slices
The “slice” syntax is a handy way to refer to sub-parts of sequences — typically strings and lists. The slice s[start:end] is the elements beginning at the start and extending up to but not including end. Suppose we have s = “Hello”
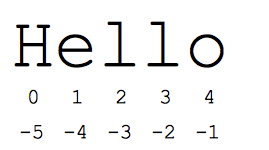
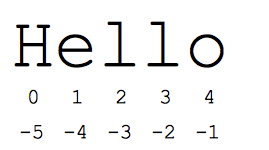
s[1:4]isell— chars starting at index 1 and extending up to but not including index 4s[1:]isello— omitting either index defaults to the start or end of the strings[:]isHello— omitting both always gives us a copy of the whole thing (this is the pythonic way to copy a sequence like a string or list)s[1:100]isello— an index that is too big is truncated down to the string length
The standard zero-based index numbers give easy access to chars near the start of the string. As an alternative, Python uses negative numbers to give easy access to the chars at the end of the string: s[-1] is the last char ‘o’, s[-2] is ‘l’ the next-to-last char, and so on. Negative index numbers count back from the end of the string:
s[-1]iso— last char (1st from the end)s[-4]ise— 4th from the ends[:-3]isHe— going up to but not including the last 3 chars.s[-3:]isllo— starting with the 3rd char from the end and extending to the end of the string.
It is a neat truism of slices that for any index n, s[:n] + s[n:] == s. This works even for n negative or out of bounds. Or put another way s[:n] and s[n:] always partition the string into two string parts, conserving all the characters. As we’ll see in the list section later, slices work with lists too.
% Format my numbers!
Python has a printf()-like the facility to put together a string. The % operator takes a printf-type format string on the left (%d is int, %s is string, %f is floating point), and the matching values in a tuple on the right (a tuple is made of values separated by commas, typically grouped inside parentheses):
text = ("%d little pigs come out or I’ll %s and %s and %s" % (3, 'huff', 'puff', 'blow down'))
print(text)
You cannot just split the line after the % as you might in other languages since by default Python treats each line as a separate statement (on the plus side, this is why we don’t need to type semi-colons on each line). To fix this, enclose the whole expression in an outer set of parenthesis — then the expression is allowed to span multiple lines. This code-across-lines technique works with the various grouping constructs detailed below: ( ), [ ], { }.
Regular Expressions – The Lasso For Data Wrangling
A regular expression is a pattern describing a subset of text. Often this concept is shortened to ‘regex’. It really was pushed forward by bioinformatics and Perl in the late 90s but has become a foundational skill. It is the Data Wrangler’s Lasso. It allows one to parse text, and scrape out just the key parts. So much of biology requires special terminology it’s a natural match.
One great thing is that it is a foundation and now a part of most languages… JavaScript, Java, VB, C #, C / C++, Python, Perl, Ruby, Delphi, R, Tcl… just to name a few. A great tool to practice Regular expressions is using Sublime3.
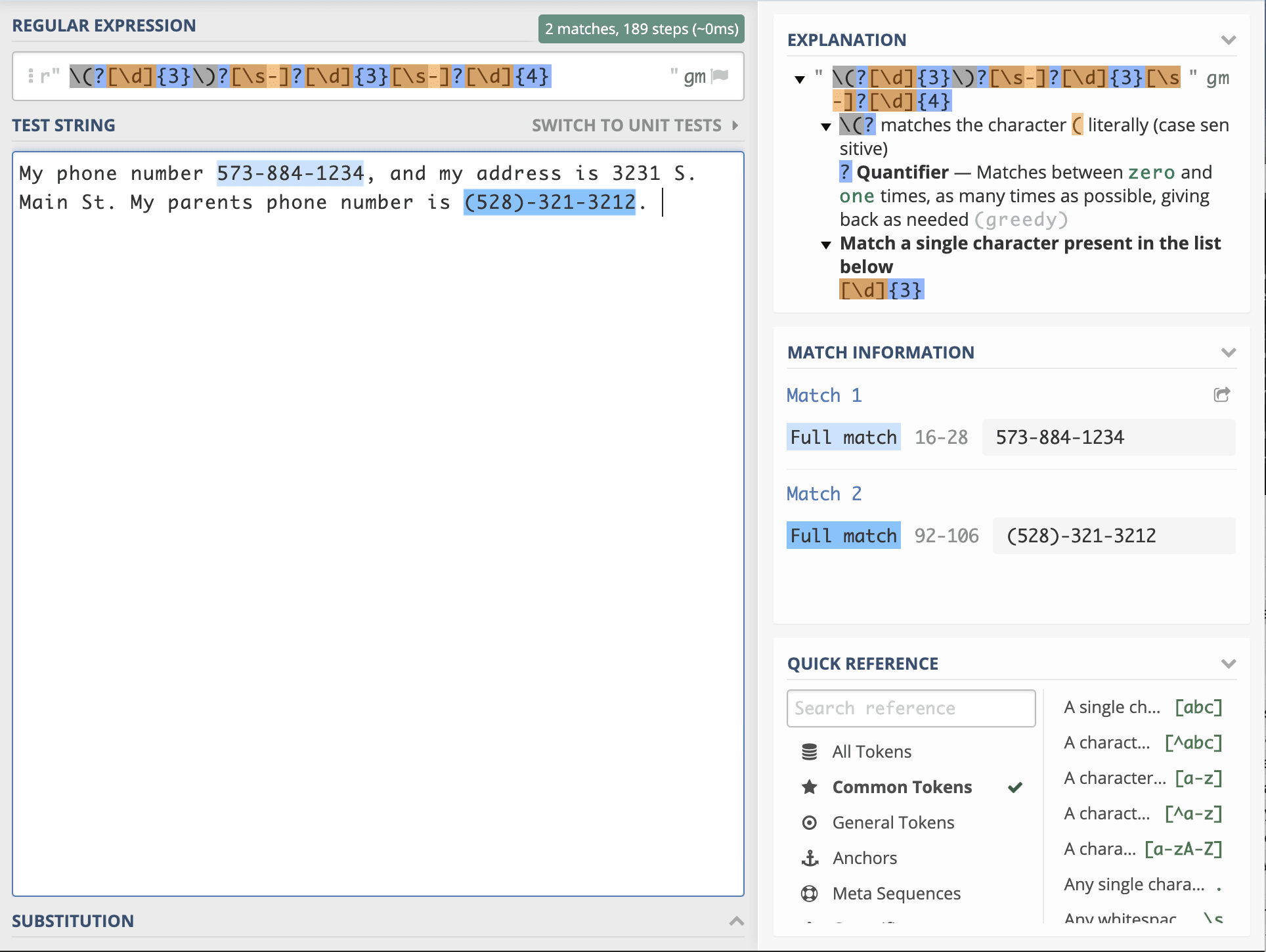

Now I’ll recommend a tool, https://regex101.com which is unbelievable effective at helping out. Let’s learn by example. Let’s say we have some text. We start by creating something to search by, and this is usually noted as being between two forward slashes. For example, /hello/ would match wherever a “hello” is found. We need to first learn some special characters which can help us match:
Regex Special characters
^The – Matches any string starting with The
end$ – Matches text/sentence that ends with end.
sometext – Matches if sometext is in the sentence
s* This is a greedy match that will match whereever 1 or more s is found
this* This will match for thi plus 0 or more ‘s’, either thissss or even thi
this+ We will match for thi plus 1 or more s ; this we will not match thi but we will match thiss This will match any
[AT]A or Twhich can also be done by (A|T)
\d Matches any digit, can be accomplished with [0-9]
\s Matches a whitespace character – tabs and spaces
. Matches any character
\D Matches anything that is not [0-9]
Regex Flags
One can put substitutions and flags in, where flags are usually at the end. We have 3 major flags:
g Match as many times as possible
m Match across multilines
i Match ignoring case
Regex: Greed is Good Greed is bad
a.+?t Matches any character as few times as possible, but at least once, provided its starts with an a and ends with a t.
a.+t Matches any character as many times as possible, but at least once, provided its starts with an a and ends with a t.
Such as as the bell toils
Greedy: Such as as the bell toils
Not Greedy: Such as as the bell toils
%%file text.txt This is content of a random set of data left by folks at (510)-919-3231. They wish you to call them at home, at their office (912)-904-2321 or at the home (323)-899-9392. But better yet you can use their 1-800-532-3233 number.
Let us say we want to search out phone numbers. The way to search a number is: \d or [0-9]. We could search for those using grepat the command-line. Note that below the symbol ! is for command-line.
!grep "\d" text.txt
This returns
This is content of a random set of data left by folks at (510)-919-3231.
Regular Expressions in Python
Regular expressions are a powerful language for matching text patterns. This page gives a basic introduction to regular expressions themselves sufficient for our Python exercises and shows how regular expressions work in Python. The Python re module provides regular expression support.
In Python a regular expression search is typically written as:
match = re.search(pat, str)
The re.search() method takes a regular expression pattern and a string and searches for that pattern within the string. If the search is successful, search() returns a match object or None otherwise. Therefore, the search is usually immediately followed by an if-statement to test if the search succeeded, as shown in the following example which searches for the pattern ‘word:’ followed by a 3 letter word (details below):
#!/usr/bin/python
import re
str = 'an example word:cat!!'
match = re.search(r'word:www', str)
# If-statement after search() tests if it succeeded
print(match)
if match:
print ('found', match.group())
else:
print ('did not find a match')
Which results in
None did not find a match
The code match = re.search(pat, str) stores the search result in a variable named “match”. Then the if-statement tests the match — if true the search succeeded and match.group() is the matching text (e.g. ‘word:cat’). Otherwise, if the match is false (None to be more specific), then the search did not succeed, and there is no matching text.
The ‘r’ at the start of the pattern string designates a python “raw” string which passes through backslashes without change which is very handy for regular expressions (Java needs this feature badly!). I recommend that you always write pattern strings with the ‘r’ just as a habit.
The basic rules of regular expression search for a pattern within a string are:
- The search proceeds through the string from start to end, stopping at the first match found
- All of the patterns must be matched, but not all of the string
- If
match = re.search(pat, str)is successful, match is not None and in particular match.group() is the matching text
Another example:
#!/usr/bin/python
import re
line = "Cats are smarter than dogs"
matchObj = re.match( r'(.*) are (.*?) .*', line, re.M|re.I)
if matchObj:
print ("matchObj.group() : ", matchObj.group())
print ("matchObj.group(1) : ", matchObj.group(1))
print ("matchObj.group(2) : ", matchObj.group(2))
else:
print ("No match!!")
This results in:
matchObj.group() : Cats are smarter than dogs matchObj.group(1) : Cats matchObj.group(2) : smarter
Another example regular expression, but using search and replace
#!/usr/bin/python
import re
phone = "2004-959-559 # This is Phone Number"
# Delete Python-style comments
num = re.sub(r'#.*$', "", phone)
print ("Phone Num : ", num)
# Remove anything other than digits
num = re.sub(r'D', "", phone)
print("Phone Num : ", num)
This yields:
Phone Num : 2004-959-559 Phone Num : 2004-959-559 # This is Phone Number
Basic Patterns
The power of regular expressions is that they can specify patterns, not just fixed characters. Here are the most basic patterns which match single chars:
- a
X9< — ordinary characters just match themselves exactly. The meta-characters which do not match themselves because they have special meanings are:. ^ $ * + ? { [ ] | ( )(details below) .(a period) — matches any single character except newline ‘n’\w— (lowercase w) matches a “word” character: a letter or digit or underbar [a-zA-Z0-9_]. Note that although “word” is the mnemonic for this, it only matches a single word char, not a whole word. W (upper case W) matches any non-word character.\b— boundary between word and non-word\s— (lowercase s) matches a single whitespace character — space, newline, return, tab, form [ nrtf]. S (upper case S) matches any non-whitespace character.\t,\n,\r— tab, newline, return\d— decimal digit [0-9] (some older regex utilities do not support but d, but they all support w and s)^= start,$= end — match the start or end of the string\inhibit the “specialness” of a character. So, for example, use.to match a period or\\to match a slash. If you are unsure if a character has special meaning, such as ‘@’, you can put a slash in front of it,\@, to make sure it is treated just as a character.
Greedy vs. Non-Greedy
This section shows a more advanced regular expression technique not needed for the exercises.
Suppose you have text with tags in it: <b>foo</b> and <i>so on</i>
Suppose you are trying to match each tag with the pattern (<.*>) — what does it match first?
The result is a little surprising, but the greedy aspect of the .* causes it to match the whole <b>foo</b> and <i>so on</i> as one big match. The problem is that the .* goes as far as is it can, instead of stopping at the first > (aka it is “greedy”).
There is an extension to regular expression where you add a ? at the end, such as .?or .+?, changing them to be non-greedy. Now they stop as soon as they can. So the pattern (<.?) will get just ‘<b>’ as the first match, and ‘</b>’ as the second match, and so on getting each <..> pair in turn. The style is typically that you use a .?, and then immediately its right look for some concrete marker (> in this case) that forces the end of the .? run.
The *? extension originated in Perl, and regular expressions that include Perl’s extensions are known as Perl Compatible Regular Expressions — pcre. Python includes pcre support. Many command-line utils etc. have a flag where they accept pcre patterns.
An older but widely used technique to code this idea of “all of these chars except stopping at X” uses the square-bracket style. For the above you could write the pattern, but instead of .* to get all the chars, use [^>]* which skips overall characters which are not > (the leading ^ “inverts” the square bracket set, so it matches any char not in the brackets).
Repetition
Things get more interesting when you use + and * to specify repetition in the pattern
+— 1 or more occurrences of the pattern to its left, e.g. ‘i+’ = one or more i’s*— 0 or more occurrences of the pattern to its left?— match 0 or 1 occurrences of the pattern to its left
Leftmost & Largest
First the search finds the leftmost match for the pattern, and second it tries to use up as much of the string as possible — i.e. + and * go as far as possible (the + and * are said to be “greedy”).
Repetition Examples
#!/usr/bin/python
import sys
import re
## i+ = one or more i's, as many as possible.
match = re.search(r'pi+', 'piiig')
print (match.group())
## Finds the first/leftmost solution, and within it drives the +
## as far as possible (aka 'leftmost and largest').
## In this example, note that it does not get to the second set of i's.
match = re.search(r'i+', 'piigiiii')
print (match.group())
## s* = zero or more whitespace chars
## Here look for 3 digits, possibly separated by whitespace.
match = re.search(r'\d\s*\d\s*\d', 'xx1 2 3xx')
if match:
print (match.group())
match = re.search(r'\d\s*\d\s*\d', 'xx12 3xx')
if match:
print (match.group())
match = re.search(r'\d\s*\d\s*\d', 'xx123xx')
if match:
print (match.group())
## ^ = matches the start of string, so this fails:
match = re.search(r'^b\w+', 'foobar')
if match:
print (match.group())
else:
print ("I failed")
## but without the ^ it succeeds:
match = re.search(r'b\w+', 'foobar')
if match:
print (match.group())
See if you can follow how this working, yielding:
piii ii 1 2 3 12 3 123 I failed bar
Emails Example
Suppose you want to find the email address inside the string xyz alice-b@google.com purple monkey. We’ll use this as a running example to demonstrate more regular expression features. Here’s an attempt
#!/usr/bin/python import sys import re str = 'purple alice-b@google.com monkey dishwasher' match = re.search(r'w+@w+', str) if match: print (match.group()) else: print(match)
The search does not get the whole email address in this case because the w does not match the ‘-‘ or ‘.’ in the address. We’ll fix this using the regular expression features below.
Square Brackets
Square brackets can be used to indicate a set of chars, so [abc] matches a or b or c. The codes w, s , etc. work inside square brackets too with the one exception that dot (.) just means a literal dot. For the emails problem, the square brackets are an easy way to add . and - to the set of chars which can appear around the @ with the pattern r'[w.-]+@[w.-]+' to get the whole email address:
#!/usr/bin/python import sys import re match = re.search(r'[w.-]+@[w.-]+', str) if match: print (match.group()) ## 'alice-b@google.com'
(More square-bracket features) You can also use a dash to indicate a range, so [a-z] matches all lowercase letters. To use a dash without indicating a range, put the dash last, e.g. [abc-]. An up-hat ^ at the start of a square-bracket set inverts it, so [^ab] means any char except a or b.
Group Extraction
The “group” feature of a regular expression allows you to pick out parts of the matching text. Suppose for the emails problem that we want to extract the username and host separately. To do this, add parenthesis ( ) around the username and host in the pattern, like this: r'([w.-]+)@([w.-]+)'. In this case, the parenthesis do not change what the pattern will match. Instead, they establish logical “groups” inside of the match text. On a successful search, match.group(1) is the match text corresponding to the 1st left parenthesis, and match.group(2) is the text corresponding to the 2nd left parenthesis. The plain match.group() is still the whole match text as usual.
#!/usr/bin/python
import sys
import re
str = 'purple alice-b@google.com monkey dishwasher'
match = re.search('([\w.-]+)@([\w.-]+)', str)
if match:
print (match.group()) ## 'alice-b@google.com' (the whole match)
print (match.group(1)) ## 'alice-b' (the username, group 1)
print (match.group(2)) ## 'google.com' (the host, group 2)
A common workflow with regular expressions is that you write a pattern for the thing you are looking for, adding parenthesis groups to extract the parts you want.
This finally shows some success:
alice-b@google.com alice-b google.com
findall
findall() is probably the single most powerful function in the re module. Above we used re.search() to find the first match for a pattern. findall() finds all the matches and returns them as a list of strings, with each string representing one match.
#!/usr/bin/python
import sys
import re
## Suppose we have a text with many email addresses
str = 'purple alice@google.com, blah monkey bob@abc.com blah dishwasher'
## Here re.findall() returns a list of all the found email strings
emails = re.findall(r'[\w.-]+@[\w.-]+', str) ## ['alice@google.com', 'bob@abc.com']
for email in emails:
# do something with each found email string
print (email)
yielding:
alice@google.com bob@abc.com
findall and Groups
The parenthesis ( ) group mechanism can be combined with findall(). If the pattern includes 2 or more parenthesis groups, then instead of returning a list of strings, findall()returns a list of tuples. Each tuple represents one match of the pattern, and inside the tuple is the group(1), group(2) .. data. So if 2 parenthesis groups are added to the email pattern, then findall() returns a list of tuples, each length 2 containing the username and host, e.g. (‘alice’, ‘google.com’).
#!/usr/bin/python
import sys
import re
f = open('test.txt', 'r')
strings = re.findall(r'\w*', f.read())
print (strings)
Once you have the list of tuples, you can loop over it to do some computation for each tuple. If the pattern includes no parenthesis, then findall() returns a list of found strings as in earlier examples. If the pattern includes a single set of parenthesis, then findall() returns a list of strings corresponding to that single group. (Obscure optional feature: Sometimes you have paren ( ) groupings in the pattern, but which you do not want to extract. In that case, write the parens with a ?: at the start, e.g. (?: ) and that left paren will not count as a group result.)
Debugging
Regular expression patterns pack a lot of meaning into just a few characters, but they are so dense, you can spend a lot of time debugging your patterns. Set up your runtime so you can run a pattern and print what it matches easily, for example by running it on a small test text and printing the result of findall(). If the pattern matches nothing, try weakening the pattern, removing parts of it so you get too many matches. When it’s matching nothing, you can’t make any progress since there’s nothing concrete to look at. Once it’s matching too much, then you can work on tightening it up incrementally to hit just what you want.
Options
The re functions take options to modify the behavior of the pattern match. The option flag is added as an extra argument to the search()or findall() etc., e.g. re.search(pat, str, re.IGNORECASE).
- IGNORECASE — ignore upper/lowercase differences for matching, so
amatches bothaandA. - DOTALL — allow dot (.) to match newline — normally it matches anything but newline. This can trip you up — you think
.*matches everything, but by default it does not go past the end of a line. Note that s (whitespace) includes newlines, so if you want to match a run of whitespace that may include a newline, you can just uses* - MULTILINE — Within a string made of many lines, allow
^and$to match the start and end of each line. Normally^/$would just match the start and end of the whole string.
Substitution
The re.sub(pat, replacement, str) function searches for all the instances of pattern in the given string, and replaces them. The replacement string can include ‘1’, ‘2’ which refer to the text from group(1), group(2), and so on from the original matching text.
Here’s an example which searches for all the email addresses, and changes them to keep the user (1) but have yo-yo-dyne.com as the host.
Datatypes: Arrays and Lists
Arrays are actually lists than be mixed datatypes, but that use numeric arrays.
#!/usr/bin/python
list = [2, 4, 7, 9]
print ("list is:",list)
list2 = [3, "test", True, 7.4]
print ("list2 is:",list2)
a = range(5) #a = [0,1,2,3,4]
print ("a:",a)
a = range(10,0,-2) #a = [10,8,6,4,2]
print ("a is now:",a)
a=[]
print ("a is now:",a)
a.append("test")
print ("a is now:",a)
a.append(5)
print ("a is now:",a)
a.pop()
print ("a is now:",a)
#-> [None, None, None, 6, None, None, None, None, None]
a.append(3)
a.append(5)
print ("a is finally and mixed:",a)
The output is:
list is: [2, 4, 7, 9] list2 is: [3, 'test', True, 7.4] a: range(0, 5) a is now: range(10, 0, -2) a is now: [] a is now: ['test'] a is now: ['test', 5] a is now: ['test'] a is finally: ['test', 3, 5]
Array functions
include len, argmax, argmin, argsort, astype, copy, info, max, mean, min, nelements, product, reshape, shape, sort,etc
#!/usr/bin/python import sys import re str = 'purple alice@google.com, blah monkey bob@abc.com blah dishwasher' ## re.sub(pat, replacement, str) -- returns new string with all replacements, ## 1 is group(1), 2 group(2) in the replacement print (re.sub(r'([w.-]+)@([w.-]+)', r'1@yo-yo-dyne.com', str))
The output is:
purple alice@google.com, blah monkey bob@abc.com blah dishwasher
Datatypes: Dictionaries
Dictionaries are defined similarly to a JSON format where each key is separated from its value by a colon (:), the items are separated by commas, and the whole thing is enclosed in curly braces. An empty dictionary without any items is {}. The values of a dictionary can be of any type, but the keys must be of an immutable data type such as strings, numbers, or tuples. (tuples are immutable lists).
#!/usr/bin/python
dict = {'PTEN': "Phosphatase and tensin homolog"}
dict['KRAS']="KRAS proto-oncogene, GTPase";
record = {'Name': 'Anna', 'Age': 8, 'Class': 'Second'}
if 'PTEN' in dict:
print (dict['PTEN']);
In this case, we now have a look-up dictionary where we can get gene:
Operators
Basic operators and math: Arithmetic: `Comparison: Logical: Exponentiation:
Others…
import os,math # Import the os and math modules
from math import * # import all functions from math
Conditionals
If, if-else, and if-elsif are common, if elif and else
#!/usr/bin/python
a=5
b=6
if (a > b):
print ("a is greater than b")
print ("blocks are defined by indentation")
elif (a < b):
print ("a is less than b")
else:
print ("a is equal to b")
a is less than b
Loops
for: for var in range(start [,stop [,inc]]): statements
Not unsimilar to IDL and basic, except for the range statement. var can be any variable. The range statement can take start and stop values, and an increment.
while: while expr: statements
Executes statements while the expression is true.
continue: continue
Skips the rest of the body of the loop for the current iteration and continue execution at the beginning of the next iteration.
break: break
Ends the execution of the current loop.
else: else
for and while loops can both have else clauses, which are executed after the loop terminates normally by falsifying the conditional, but else clauses are not executed when a loop terminates via a break statement.
foreach: for x in array: statements
Loops over the array given by array. On each iteration, the value of the current element is assigned to x and the internal array pointer is advanced by one.
value = "I am some text"
for j in range(10):
print ("Value number " + str(j) +" is "+value[j])
x=0
for j in range(10,0,-2):
x = x + j
print ('x is',x)
b=1
a=15
while (b < a):
print ("b ",b," is less than a,",a,".")
b=b+1
k=0
for j in range(0,10):
while(k < j):
print ("j = " + str(j) + " k = "+str(k))
if (j == 1): break
k=k+1
print ("j equals k or j equals 1")
a = ["abc","def","ghi"]
for x in a:
print ('x is ',x)
Value number 0 is I Value number 1 is Value number 2 is a Value number 3 is m Value number 4 is Value number 5 is s Value number 6 is o Value number 7 is m Value number 8 is e Value number 9 is x is 10 x is 18 x is 24 x is 28 x is 30 b 1 is less than a, 15 .
Functions
Functions are defined following several conventions, noting that variables are private unless otherwise stated. Functions in Python are defined like this:
def double(n):
n = n * 2
return n
x = 2
x = double(x)
print ('x:',x)
def repeat(s, exclaim):
"""
Returns the string 's' repeated 3 times.
If exclaim is true, add exclamation marks.
"""
result = s + s + s # can also use "s * 3" which is faster (Why?)
if exclaim:
result = result + '!!!'
return result
print(repeat('s',1))
print(repeat('s',0))
Output is:
sss!!! sss
Notice also how the lines that make up the function or if-statement are grouped by all having the same level of indentation. We also presented 2 different ways to repeat strings, using the + operator which is more user-friendly, but * also works because it’s Python’s “repeat” operator, meaning that '-' * 10 gives '----------', a neat way to create an onscreen “line.” In the code comment, we hinted that * works faster than +, the reason being that * calculates the size of the resulting object once whereas with +, that calculation is made each time + is called. Both + and * are called “overloaded” operators because they mean different things for numbers vs. for strings (and other data types).
The def keyword defines the function with its parameters within parentheses and its code indented. The first line of a function can be a documentation string (“docstring”) that describes what the function does. The docstring can be a single line, or a multi-line description as in the example above. (Yes, those are “triple quotes,” a feature unique to Python!) Variables defined in the function are local to that function, so the “result” in the above function is separate from a “result” variable in another function. The return statement can take an argument, in which case that is the value returned to the caller.
#!/usr/bin/python
# Defines a "repeat" function that takes 2 arguments.
def repeat(s, exclaim):
"""
Returns the string 's' repeated 3 times.
If exclaim is true, add exclamation marks.
"""
result = s + s + s # can also use "s * 3" which is faster (Why?)
if exclaim:
result = result + '!!!'
return result
def main():
print (repeat('Yay', False)) ## YayYayYay
if __name__ == '__main__':
main()
Here is code that calls the above repeat() function, printing what it returns:
YayYayYay
When running, functions must be defined by the execution of a def before they are called. It’s typical to def a main() function towards the bottom of the file with the functions it calls above it.
File i/o examples
Lets create a few test files:
file1:
%%file lookup.csv Date,name 123,baby abc,cat
File 2:
%%file test.csv Date,Foo,Bar 123,456,789 abc,def,ghi
Creating a script called io.py
A lookup script to substitute the first column of a csv file with a dictionary csv file from another file.
%%file io.py
#!/usr/bin/python
import sys
import re
with open(sys.argv[1], 'r') as f:
for line in f:
line = re.sub(r'<.*?>', "", line) # Regex
line=line.rstrip() # strips the line break
if len(line) > 0: # insures that there is some text in line
print(line)
Now we run it:
!python io.py lookup.csv test.csv
./io.py lookup.csv test.csv ['name', 'Foo', 'Bar'] ['baby', '456', '789'] ['cat', 'def', 'ghi']
More on Modules and their Namespaces
Suppose you’ve got a module binky.py which contains a function def foo(). The fully qualified name of that foo function is binky.foo. In this way, various Python modules can name their functions and variables whatever they want, and the variable names won’t conflict — module1.foo is different from module2.foo. In the Python vocabulary, we’d say that binky, module1, and module2 each have their own “namespaces,” which as you can guess are variable name-to-object bindings.
For example, we have the standard sys module that contains some standard system facilities, like the argv list, and exit() function. With the statement import sys you can then access the definitions in the sys module and make them available by their fully-qualified name, e.g. sys.exit(). (Yes, sys has a namespace too!)
def main():
print repeat('Yay', False) ## YayYayYay
print repeat('Woo Hoo', True) ## Woo HooWoo HooWoo Hoo!!!
There is another import form that looks like this: from sys import argv, exit. That makes argv and exit() available by their short names; however, we recommend the original form with the fully-qualified names because it’s a lot easier to determine where a function or attribute came from.
There are many modules and packages which are bundled with a standard installation of the Python interpreter, so you don’t have to do anything extra to use them. These are collectively known as the “Python Standard Library.” Commonly used modules/packages include:
sys— access to exit(), argv, stdin, stdout, …re— regular expressionsos— operating system interface, file system
Learn through an Exercise
Setup
We have two files, and a problem we need to solve. One file is a list of genes and their expression level produced from a program called Feature Counts. The gene names use their Ensembl gene name – not helpful as ENSGXXX just isn’t meaningful to the biologist. The file is already available and ready to look at it in a common directory. This is shown as gene_counts.csv and the first several lines are below. It can be retrieved at https://github.com/davcraig75/tgrn510.
Lets do this using command-line. The command wget will download the file that is at this address:
!wget https://github.com/davcraig75/tgrn510/raw/master/gene_counts.csv.gz
HTTP request sent, awaiting response... 200 OK Length: 302231 (295K) [application/octet-stream] Saving to: ‘gene_counts.csv.gz’ gene_counts.csv.gz 100%[===================>] 295.15K --.-KB/s in 0.04s 2020-04-17 04:58:02 (6.69 MB/s) - ‘gene_counts.csv.gz’ saved [302231/302231]
This file is compressed, we need to uncompress it. When we do the ending of .gz will be removed. It will then be readable using a regular text reader, and we will use head.
!gunzip gene_counts.csv.gz !ls
We can look at the first few lines to make sure we understand it:
!head gene_counts.csv
There are 2 columns and it is comma delimited.
Name,1009_POOL
ENSG00000000003.14,2.653706
ENSG00000000005.5,0
ENSG00000000419.12,20.734869
ENSG00000000457.13,11.17041
ENSG00000000460.16,6.406945
ENSG00000000938.12,146.736241
ENSG00000000971.15,2.170721
ENSG00000001036.13,8.940068
ENSG00000001084.12,18.911549
The goal is to substitute for the canonical gene name using the Ensembl gene definition files, effectively replacing column 1.
The second file is the GTF file, and it has a standard definition, described here and also shown below (Homo_sapiens.GRCh37.75.gtf). The location is here, so we will use wget from a server.
!wget --no-check-certificate https://itg.usc.edu/site/Homo_sapiens.GRCh37.75.gtf.gz !gunzip Homo_sapiens.GRCh37.75.gtf.gz !ls
Connecting to itg.usc.edu (itg.usc.edu)|128.125.215.226|:443... connected. WARNING: cannot verify itg.usc.edu's certificate, issued by ‘CN=InCommon RSA Server CA,OU=InCommon,O=Internet2,L=Ann Arbor,ST=MI,C=US’: Unable to locally verify the issuer's authority. HTTP request sent, awaiting response... 200 OK Length: 39344043 (38M) [application/x-gzip] Saving to: ‘Homo_sapiens.GRCh37.75.gtf.gz’ Homo_sapiens.GRCh37 100%[===================>] 37.52M 18.8MB/s in 2.0s 2020-04-17 05:35:28 (18.8 MB/s) - ‘Homo_sapiens.GRCh37.75.gtf.gz’ saved [39344043/39344043]
We have unzipped file. Lets look at the first few lines in the command-line:
!head Homo_sapiens.GRCh37.75.gtf
#!genome-build GRCh37.p13 #!genome-version GRCh37 #!genome-date 2009-02 #!genome-build-accession NCBI:GCA_000001405.14 #!genebuild-last-updated 2013-09 1 pseudogene gene 11869 14412 . + . gene_id "ENSG00000223972"; gene_name "DDX11L1"; gene_source "ensembl_havana"; gene_biotype "pseudogene"; 1 processed_transcript transcript 11869 14409 . + . gene_id "ENSG00000223972"; transcript_id "ENST00000456328"; gene_name "DDX11L1"; gene_source "ensembl_havana"; gene_biotype "pseudogene"; transcript_name "DDX11L1-002"; transcript_source "havana"; 1 processed_transcript exon 11869 12227 . + . gene_id "ENSG00000223972"; transcript_id "ENST00000456328"; exon_number "1"; gene_name "DDX11L1"; gene_source "ensembl_havana"; gene_biotype "pseudogene"; transcript_name "DDX11L1-002"; transcript_source "havana"; exon_id "ENSE00002234944"; 1 processed_transcript exon 12613 12721 . + . gene_id "ENSG00000223972"; transcript_id "ENST00000456328"; exon_number "2"; gene_name "DDX11L1"; gene_source "ensembl_havana"; gene_biotype "pseudogene"; transcript_name "DDX11L1-002"; transcript_source "havana"; exon_id "ENSE00003582793"; 1 processed_transcript exon 13221 14409 . + . gene_id "ENSG00000223972"; transcript_id "ENST00000456328"; exon_number "3"; gene_name "DDX11L1"; gene_source "ensembl_havana"; gene_biotype "pseudogene"; transcript_name "DDX11L1-002"; transcript_source "havana"; exon_id "ENSE00002312635";
If look carefully, we can see it defines much more including lines for indicating exons and other features. What we want is the “gene_name” for those lines that are designated as a gene in the 3rd column. We need to create a “Lookup Dictionary” for gene_id, such that for example if we saw ENSG00000223972, we would replace it with DDX11L1.
Step 1: Getting command line arguments, importing some functions, variables
We want to get some information from the command-line, allowing us to pass the name of the Salmon script just in case we want to use this same script again. Since this will change, lets make sure and give the passed information its own variable name
%%file gene2ensemble.py import sys # Load in module that accesses the command line my_dict_file= sys.argv[1] # Assign my_file to whatever was argument 1
Step 2: Loops to the rescue
Lets create a loop to go through the file, line-by-line and just to make sure things work, lets test it printing the line.
%%file gene2ensemble.py
import sys # Load in module that accesses the command line
import fileinput # This module gives us the ability to read files
my_dict_file= sys.argv[1] # Assign my_file to whatever was argument 1
# Using the newly added fileinput library, we have it read the file line by line, assigning to each_line_of_text variable
for each_line_of_text in fileinput.input(my_dict_file):
print (each_line_of_text)
In this, we use a module/library called fileinput, and we use a function to read the input and loop through the body of our file, line by line, and each time assigning it to each_line_of_text variable. Then we do something simple and print each_line_of_text. Now lets it, just looking at top few lines.
!python gene2ensemble.py Homo_sapiens.GRCh37.75.gtf | head
#!genome-build GRCh37.p13
#!genome-version GRCh37
#!genome-date 2009-02
#!genome-build-accession NCBI:GCA_000001405.14
#!genebuild-last-updated 2013-09
Traceback (most recent call last):
File "./test.py", line 8, in <module>
print each_line_of_text
IOError: [Errno 32] Broken pipe
The Broken pipe is an error because we used head, which we can ignore or we can google how to address that. For now, I’m not addressing it.
Step 3: Simple regex matching & a conditional
In this file, I want to build a dictionary since I can map the gene_id to gene_name, creating a dictionary. The file though contains a lot of information, and as I mentioned, I really only need those lines where the 3rd column is gene. So I load in the regular expression library which comes with a few really great functions including search, match, and some others that can be used with standard regular expression prose. in this case, I match on those where I create a conditional on the third set of characters following a tab is gene. If this conditional is meant I then print the line.
%%file gene2ensemble.py
import sys # Load in module that accesses the command line
import fileinput # This module gives us the ability to read files
import re # This imports the regex capability
my_file= sys.argv[1] # Assign my_file to whatever was argument 1
for each_line_of_text in fileinput.input(my_file):
if re.match(r'.*\t.*\tgene\t', each_line_of_text):
print (each_line_of_text)
By now we should be readily observing the indenting – once to indicate the text that is within the loop, and one to indicate those things require a condition be met (e.g. the “if”).
Step 4: Arrays come alive with regex
Now, I want to pull out some text using regex and a complex match from this sentence as an example:
1 pseudogene gene 11869 14412 . + . gene_id "ENSG00000223972"; gene_name "DDX11L1"; ge...
To figure out the regular expression, I use regex101.com – that’s just me – where I put an example line, and I start composing the matching regular expression. I remember that . is any character, .* mean 0 or more of the previous (any character) to the maximum match (greedy matching, and that .*? is a non greedy match. I can match thus where I look for gene_id, followed by a space, and " (literal, so I add a backslash), a series of characters matching untill I see another ". At this point, I’m at gene_id \".*?\" . However, I need to pull out text and not all of that match. Now I don’t need the full match, I actually just want the text between the two quotes. I can capture some of the text by using parenthesis, and thus I add a parenthesis, so gene_id \"(.*?)\" will give me a match on just what is between the parenthesis.
%%file gene2ensemble.py
import sys # Load in module that accesses the command line
import fileinput # This module gives us the ability to read files
import re # This imports the regex capability
my_file= sys.argv[1] # Assign my_file to whatever was argument 1
for each_line_of_text in fileinput.input(my_file):
if re.match(r'.*\t.*\tgene\t', each_line_of_text):
gene_id_matches = re.findall('gene_id \"(.*?)\";',each_line_of_text)
gene_name_matches = re.findall('gene_name \"(.*?)\";',each_line_of_text)
print ('gene_id is ' + str(gene_id_matches))
print ('gene_name is ' + str(gene_name_matches))
print ('gene_id[0] is ' + gene_id_matches[0])
So I have made some matches, and they are going to two variables. When I print these variables, gene_id_matches and gene_name_matches , I’ll see that they are actually arrays. I test this out I’ll see:
!python gene2ensemble.py Homo_sapiens.GRCh37.75.gtf | head
['ENSG00000223972']
['DDX11L1']
gene_id[0] is ENSG00000223972
['ENSG00000227232']
['WASH7P']
gene_id[0] is ENSG00000227232
['ENSG00000243485']
['MIR1302-10']
gene_id[0] is ENSG00000243485
['ENSG00000237613']
Step 5: Now… Storing in a dictionary
A tremendous use of associative arrays, dictionaries, or hashes (names that largely refer to data stores which are key/value) is as lookup vehicles. Essentially, we want a variable called Lookup_geneID, which when given a key of a gene_ID, yields the gene_name. Thus print Lookup_geneID[‘ENSG1232343’] would give BRAGG (if that were a real gene).
So simply, I initialize a dictionary at step one, that I will slowly fill as I traverse the file. Now to test it, I need to spit out the contents. Since I am new to python, I’ll just google how to print out a JSON. This is my sanity check at this step.
%%file gene2ensemble.py
import sys # Load in module that accesses the command line
import fileinput # This module gives us the ability to read files
import re # This imports the regex capability
my_file= sys.argv[1] # Assign my_file to whatever was argument 1
Lookup_geneID={}
for each_line_of_text in fileinput.input(my_file):
if re.match(r'.*\t.*\tgene\t', each_line_of_text):
gene_id_matches = re.findall('gene_id \"(.*?)\";',each_line_of_text)
gene_name_matches = re.findall('gene_name \"(.*?)\";',each_line_of_text)
if gene_id_matches:
if gene_name_matches:
Lookup_geneID[gene_id_matches[0]] = gene_name_matches[0]
print ("Entry:" + gene_id_matches[0] + " key is " + gene_name_matches[0])
We can see our results:
!python gene2ensemble.py Homo_sapiens.GRCh37.75.gtf | head
I just created an entry where ENSG00000223972 is a key and its value is DDX11L1
I just created an entry where ENSG00000227232 is a key and its value is WASH7P
I just created an entry where ENSG00000243485 is a key and its value is MIR1302-10
I just created an entry where ENSG00000237613 is a key and its value is FAM138A
I just created an entry where ENSG00000268020 is a key and its value is OR4G4P
I just created an entry where ENSG00000240361 is a key and its value is OR4G11P
I just created an entry where ENSG00000186092 is a key and its value is OR4F5
I just created an entry where ENSG00000238009 is a key and its value is RP11-34P13.7
I just created an entry where ENSG00000239945 is a key and its value is RP11-34P13.8
I just created an entry where ENSG00000233750 is a key and its value is CICP27
Traceback (most recent call last):
File "gene2ensemble.py", line 13, in <module>
print ("I just created an entry where " + gene_id_matches[0] + " is a key and its value is " + gene_name_matches[0])
BrokenPipeError: [Errno 32] Broken pipe
Ignore the error since we know our command in BASH caused it by piping to head
Step 6. Reading a second file, splitting the input into an array
We have created our dictionary, and now we want to read our file that needs to be changed. We will have this be the second argument we get, and we will assign it to a variable input_file_to_change , then loop through line by line and split up the results into an array based on tabs. An array is reasonable, particularly in a tsv file, and split using regex is a nice way to split up a row, and place the value in each column within the array.
%%file gene2ensemble.py
import sys # Load in module that accesses the command line
import fileinput # This module gives us the ability to read files
import re # This imports the regex capability
my_file= sys.argv[1] # Assign my_file to whatever was argument 1
Lookup_geneID={}
for each_line_of_text in fileinput.input(my_file):
each_line_of_text = each_line_of_text.strip('\n')
if re.match(r'.*\t.*\tgene\t', each_line_of_text):
gene_id_matches = re.findall('gene_id \"(.*?)\";',each_line_of_text)
gene_name_matches = re.findall('gene_name \"(.*?)\";',each_line_of_text)
if gene_id_matches:
if gene_name_matches:
Lookup_geneID[gene_id_matches[0]] = gene_name_matches[0]
# Read in the file we which to modify
input_file_to_change=sys.argv[2]
for each_line_of_text in fileinput.input(input_file_to_change):
splitcolumn_array = re.split(',',each_line_of_text.replace( '"' ,'').replace('\n',''))
print(splitcolumn_array)
This file will take a bit longer, because matching can be slow. As a note, its possible to make regex faster by predesignating the ‘regex’ expression, that is to use the compile command.
Step 7. Hey – how do we check to see if the term is in the dictionary?
We might guess – or we might find by way of error, that we need to actually have a key in our dictionary in order to get the value. How does one check to see if a key exists in our dictionary? The answer isn’t in our primer, so we do the search in google, where we see the examples of if text_in_columns[0] in Lookup_geneID: Alas we have conditional to check to make sure its a valid key. If so, we can then substitute and print, finishing our program first draft.
We add a few lines to fix the header…
%%file gene2ensemble.py
import sys # Load in module that accesses the command line
import fileinput # This module gives us the ability to read files
import re # This imports the regex capability
my_file= sys.argv[1] # Assign my_file to whatever was argument 1
Lookup_geneID={}
for each_line_of_text in fileinput.input(my_file):
each_line_of_text = each_line_of_text.strip('\n')
if re.match(r'.*\t.*\tgene\t', each_line_of_text):
gene_id_matches = re.findall('gene_id \"(.*?)\";',each_line_of_text)
gene_name_matches = re.findall('gene_name \"(.*?)\";',each_line_of_text)
if gene_id_matches:
if gene_name_matches:
Lookup_geneID[gene_id_matches[0]] = gene_name_matches[0]
# Read in the file we which to modify
input_file_to_change=sys.argv[2]
for each_line_of_text in fileinput.input(input_file_to_change):
splitcolumn_array = re.split(',',each_line_of_text.replace( '"' ,'').replace('\n',''))
if splitcolumn_array[0] in Lookup_geneID:
print (Lookup_geneID[splitcolumn_array[0]] + "\t" + splitcolumn_array[1])
Lets run, and redirect into a new file.
TSPAN6 0.0334633 TNMD 0 DPM1 1.86011 SCYL3 1.5432 C1orf112 1.198 FGR 20.029 CFH 0.392603 FUCA2 1.02429 GCLC 3.14715 NFYA 2.15042
It works! In this process, we’ve learned associative arrays, arrays, regex, loops, conditionals, functions, variables and most of our key concepts we needed for basic python.

